The newly sequenced genome of a dainty, quill-like sea creature called a lancelet provides the best evidence yet that vertebrates evolved over the past 550 million years through a four-fold duplication of the genes of more primitive ancestors.
The late geneticist Susumu Ohno argued in 1970 that gene duplication was the most important force in the evolution of higher organisms, and Ohno's theory was the basis for original estimates that the human genome must contain up to 100,000 distinct genes.
Instead, the Human Genome Project found that humans today have only 20,000 to 25,000 genes, which means that, if our ancestors' primitive genome doubled and redoubled, most of the duplicate copies of genes must have been lost. An analysis of the lancelet, or amphioxus, genome, published in the June 19 issue of Nature, shows this to be the case.
"Amphioxus and humans had a common ancestor 550 million years ago, which allows us to use amphioxus as a surrogate for that ancestor in terms of understanding how vertebrate genomes evolved," said Daniel S. Rokhsar, a faculty member in the University of California, Berkeley's Center for Integrative Genomics and program head for computational genomics at the Department of Energy Joint Genome Institute (JGI) in Walnut Creek, Calif. Rokhsar and JGI post-doctoral fellow Nicholas H. Putnam performed the sequencing, assembly and genome-wide analyses of the amphioxus genome and are lead authors of the Nature paper.
"If you compare the 23 chromosomes of humans with the 19 chromosomes of amphioxus, you find that both genomes can be expressed in terms of 17 ancestral pieces. So, we can say with some confidence that 550 million years ago, the common ancestor of amphioxus and humans had 17 chromosomal elements."
JGI post-doctoral fellow Nicholas H. Putnam, along with Rokhsar and a large international group of collaborators from the United States, Japan, the United Kingdom, and Spain, reconstructed what happened next. Each of those 17 ancestral segments was duplicated twice in the evolution of vertebrates, after which most of the routine "housekeeping" genes lost the extra copies. Those left, totaling a couple thousand genes, found new functions that, Putnam said, make us different from all other creatures.
"These few thousand genes have been retooled to make humans more elaborate than their simpler ancestors. They are involved in setting up the body plan of an animal and differentiating different parts of the animal," he said. "The hypothesis, pretty strongly supported by this data, is that the multiplication of this particular kind of gene and differentiation into different functions was important in the formation of vertebrates as we know them."
"The most exciting thing that the amphioxus genome does is provide excellent evidence for the idea that Ono proposed in 1970, that the human genome had undergone two rounds of whole-genome duplication with subsequent losses," said coauthor Linda Z. Holland, an expert on the biology and genetics of amphioxus at the Scripps Institution of Oceanography at UC San Diego who led the community effort to sequence and annotate the genome. "We have been kicking that idea around with very little proof for a long time. This genome sequence really clinches that."
Interestingly, the sea squirt Ciona intestinalis, a tunicate, was previously thought to belong to the the earliest chordate lineage because of the sea squirt's very simple body plan. Comparison of the lancelet, sea squirt and human genomes, however, show instead that the lancelet lineage diverged before the tunicates and vertebrates.
"Sea squirts and their relatives have taken the basic chordate genome and simplified it in various ways, while amphioxus retains those features in its genome," said Rokhsar.
The researchers are trying to reconstruct what happened at the end of the Cambrian period 550 million years ago, when a creature similar to the lancelet evolved and diverged into three types of chordates: cephalocordates like the lancelet; urochordates like the sea squirt; and vertebrates like us. Cephalocordates and urochordates are invertebrates that have a flexible notochord rather than a bony spine protecting their spinal cord.
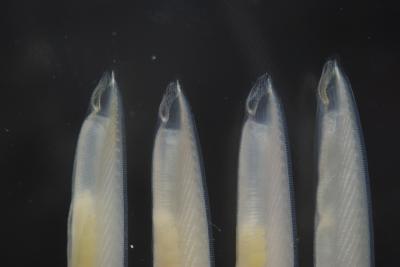
The Florida lancelet, Branchiostoma floridae, looks a lot like a fish, but has a stiff cartilaginous notochord that it vibrates to swim and burrow in the sand, where it nestles during the day with its head sticking out to filter phytoplankton from the water. The lancelets typically emerge from the sand at night and swim with lateral movements.
"If you think about what the original ancestor of all bilaterally symmetric animals looked like, you can make a pretty good argument that it was a sort of worm-like creature - a finger length-long, floppy kind of worm like amphioxus," Rokhsar said.
With the aim of understanding the evolution of chordates, the JGI sequenced the amphioxus genome using lancelets that Holland collected from the 15-foot-deep waters of Tampa Bay, Fla. A thorough analysis led by Putnam, Rokhsar, Holland and colleagues Peter Holland of Oxford University in the U.K. and Noriyuki Satoh of Kyoto University shows that the creature's 19 chromosomes map onto the human genome in 17 segments, each of which is represented four times in the human genome.
"The human genome is a mosaic of these 17 ancestral pieces constructed by two rounds of duplication, followed by gene loss and chromosome rearrangments and fusions. That took some computational gymnastics to sort out, but the evidence is still there," said Rokhsar.
Putnam's analysis of the genome shows that many of the duplicated genes were subsequently lost, "mainly the housekeeping genes that code for structural proteins, enzymes, metabolic pathways - things you don't really need more than one copy of," Linda Holland said.
However, Rokhsar added, "a class of a couple thousand genes did not return to a single copy. Those extra copies of genes acquired some function that prevented them from being lost. The vast majority are regulatory genes highly enriched for transcription factors and genes involved in developmental signaling."
Holland, a research biologist at Scripps in the Marine Biology Research Division, is the lead author of a companion paper appearing this week in the July issue of the journal Genome Research that looks at these genes in detail to see how vertebrates have employed old genes for new functions.
"We are finding that today's complicated vertebrate has not invented a lot of new genes to become complicated," she said. "Amphioxus shows us that vertebrates have taken old genes and recombined them, changed their regulation and perhaps changed the gene function."
Such duplication has given humans and other vertebrates a much larger "toolkit" for making various structures that are absent in amphioxus, including cells for pigment and collagen type II-based cartilage, for example.
Putnam noted another interesting finding reinforced by the amphioxus genome: Most creatures have a lot more genetic variation than humans. While two humans typically differ at only one nucleic acid per thousand in the genome, two lancelets differ at one of every 16 nucleic acids.
"Marine invertebrates actually vary about 6 percent, which means that, on average, one of every 16 bases is different, which is pretty remarkable - it's the difference between humans and certain types of apes," Putnam said. "Humans really are a special case, because of the recent out-of-Africa bottleneck and because of the size of our population. There is a lot less variation than in these little wormy guys that live by the millions in shallow water."
Coauthors include Takeshi Kawashima, formerly of UC Berkeley's Center for Integrative Genomics; Uffe Hellsten, Astrid Terry, Inna Dubchak, Igor V. Grigoriev, Erika Lindquist, Susan Lucas, Len A. Pennacchio and Asaf A. Salamov of the JGI; Thomas Butts, Rebecca F. Furlong and Peter W. H. Holland of the University of Oxford in the United Kingdom; David E. K. Ferrier of the University of St. Andrews in the U.K.; Marc Robinson-Rechavi of the University of Lausanne, Switzerland; Eiichi Shoguchi, Yutaka Satou and Nori Satoh of Kyoto University, Japan; Jr-Kai Yu, Tatjana Sauka-Spengler and Marianne Bronner-Fraser of the California Institute of Technology; Èlia Benito-Gutiérrez of the U.K.'s National Institute for Medical Research; Jordi Garcia-Fernàndez of the University of Barcelona, Spain; Jeremy J. Gibson-Brown and Amy C. Horton of Washington University in St. Louis, Mo.; Pieter J. de Jong and Kazutoyo Osoegawa of Children's Hospital Oakland Research Institute in Oakland, Calif.; Jerzy Jurka and Vladimir Kapitonov of the Genetic Information Research Institute in Mountain View, Calif.; Yuji Kohara and Tadasu Shin-I of Japan's National Institute of Genetics; Yoko Kuroki and Atsushi Toyoda of the RIKEN Genomic Sciences Center in Yokohama, Japan; Jeremy Schmutz of the Stanford Human Genome Center in Palo Alto, Calif.; and Asao Fujiyama of the National Institute of Informatics in Tokyo, Japan.
The amphioxus genome was sequenced by the Department of Energy (DOE) Joint Genome Institute under the auspices of the DOE's Office of Science, Biological and Environmental Research Program, Lawrence Berkeley National Laboratory, Lawrence Livermore National Laboratory and Los Alamos National Laboratory. UC Berkeley's Center for Integrative Genomics is funded by the Gordon and Betty Moore Foundation.





Comments