It may take just one or two proteins to polish off a simple cellular task, but life-or-death matters, such as caring for the ends of chromosomes known as telomeres, require interacting crews of proteins, all with a common goal but each with a specialized task.
Researchers at the Salk Institute for Biological Studies led by Vicki Lundblad, Ph.D., a Professor in the Molecular and Cell Biology Laboratory, have discovered that a protein that helps elongate chromosome ends—and hence saves cells from premature growth arrest—likely recognizes where to report to work through a common fold. Those findings are reported in the online edition of Nature Structure and Molecular Biology.
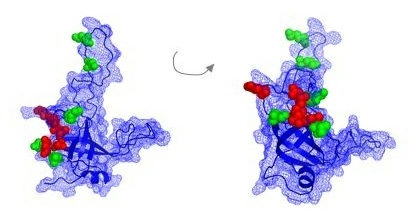
“Our studies have predicted a specific 3-dimensional shape displayed by a key protein that interacts with telomeres,” said Lundblad, explaining that analysis of this protein, called Est3, had been up to now particularly problematic. “This prediction has now provided us with a crucial tool that has propelled forward our studies of this protein.”
When cells divide, telomeres – the tails of repetitive DNA that extend from the ends of each chromosome – become progressively shorter. Telomeres function like cellular clocks: as they become critically stubby cells are no longer capable of dividing and undergo growth arrest termed cellular “senescence”. Fortunately, over the course of a cell’s lifetime, an enzyme complex known as telomerase works to restore telomeres to a more youthful length after each cell division.
Although the precise structure of human telomerase is not yet known, Lundblad had defined its budding yeast equivalent as containing three Est proteins complexed to an RNA ribbon: Est2 and the RNA do the heavy catalytic lifting in terms of telomere-reconstruction, while Est1 and Est3 orchestrate the process. But just how the two proteins ensured that things run smoothly was still unclear.
An earlier study in Lundblad’s group already answered this question for Est1: they showed that Est drags the telomerase complex to telomeres, an activity that is required to keep yeast cells continuously dividing. “Without Est1, telomerase cannot get to the ends of chromosomes, and thus telomeres shorten,” explained Lundblad.
In the current study Lundblad and co-first authors post-doctoral researcher Jaesung Lee, Ph.D., and graduate student Edward Mandell turned their attention to Est3 by first examining its computer-generated three-dimensional structure. They found that, like several other telomeric proteins, Est3 exhibited an architectural element known as an “OB” fold. Using computer models to scrutinize the fold surface, the group predicted that specific amino acids on one face of the fold might be required for interaction with the telomerase complex.
Laborious benchwork confirmed their predictions: the group biochemically inactivated each candidate amino acid separately, put each of those mutant proteins back into yeast cells, and then monitored whether that manipulation had any effect on telomere length or cell survival.
Their exhaustive analysis proved highly fruitful in terms of telomerase regulation: inactivation of a handful of suspect amino acids on one side of the surface of the protein produced shorter telomeres, showing that telomerase was now severely impaired.
Alterations in one cluster of amino acids lead to failure of Est3 to interact with its other Est protein partners. Disrupting a second set of amino acids leaves the telomerase complex intact, but with an inactive Est3 protein. Addressing what makes Est3 inactive is the subject of ongoing investigation.
“The recent increase in the ability to predict protein structure is now permitting detailed analysis of many proteins, including proteins like Est3, whose function had been mysterious,” says Lundblad.
An unanticipated but highly intriguing finding came from comparison of Est3 structure to that of other proteins. The group found that Est3 highly resembles a mammalian telomere-associated protein called TPP1. “This is surprising because TPP1 is not a subunit of the telomerase complex,” explained Lundblad, noting that instead TPP1’s job is to bind to and shelter telomeres from cellular repair enzymes that might mistake shaggy chromosome ends for damaged DNA — not to regulate their construction.
Factors that regulate telomerase activity are a very hot topic in biomedicine: sluggish telomerase activity promotes premature cell death and may underlie diseases of aging via telomere shortening, while hyperactive telomerase could promote uncontrolled cell division and cellular immortality associated with cancer.
The common architecture of telomere-associated proteins that serve discrete functions is further evidence that shared protein motifs, as exemplified by the signature OB-fold, do not always determine what biochemical task a protein accomplishes but rather signal where it does it.
Or as Lundblad concluded, “What we now propose that this particular protein fold, which both Est3 and TPP1 exhibit, is uniquely suited to the telomere.”
Researchers who also contributed to the study include graduate student Timothy Tucey and Danna Morris, a former graduate student in Lundblad’s lab.
The research was supported by funding from the National Institute on Aging at the NIH.






Comments