In the past 30 years, food allergies have become increasingly common in the United States. Changes to human genetics can’t explain the sudden rise. That is because it takes many generations for changes to spread that widely within a population. Perhaps the explanation lies in changes to our environment, particularly our internal environment.
Shifting lifestyle practices over the last half-century – increasing antibiotic and antimicrobial use, surface sterilization, air filtration and changes to diet – have changed our internal environment and wiped out important bacteria with beneficial health effects.
For many years, my research group at the University of Chicago has been exploring the role that intestinal bacteria play in preventing allergic responses to food. Bacteria, together with viruses, fungi and other small organisms living in and on our bodies, collectively make up the microbiome and play a critical supporting role in health.
The microbiome is our internal environment. Humans and microbes have “grown up” together: As humans evolved, so did their microbes. We tend to think of health practices as changing slowly, but from the perspective of the bacteria in our guts, changes in their composition and function have happened more quickly – and the results are dramatic.
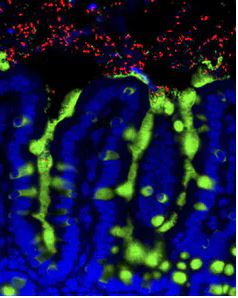
Bacteria (red) reside among the epithelial cells (blue) and the mucus (green) of a mouse small intestine.UChicago, cnagler@bsd.uchicago.edu, CC BY-SA
Intestinal bacteria and allergies
Several years ago, my research group, together with a collaborator in Italy, Roberto Berni Canani, was comparing the bacteria present in infants with a diagnosed cow’s milk allergy to those without. We found some remarkable differences between the two groups. This led us to wonder whether the different bacteria present in each of the two groups are sufficient to protect against allergy. And if so, could we figure out why?
To test this idea, we transplanted the entire microbiome of the two different groups – the healthy infants and those allergic to cow’s milk – into special laboratory mice that were bred in a completely sterile environment, with no bacteria of their own. The idea was simple: If we transplant the different groups of bacteria into mice, will the mouse become allergic to cow’s milk or not?
When we did this, we were stunned by the results: The bacteria from a healthy infant could protect the mouse from developing an anaphylactic response to a cow’s milk protein, while the bacteria from a cow’s milk allergic infant did not.
A new diagnostic?
When we cataloged bacteria present in the mice colonized with healthy bacteria and those present in the mice colonized with the cow’s milk allergic bacteria, we were able to calculate a ratio of protective to nonprotective groups. This ratio could accurately predict whether or not the infants had an allergy. We also learned that the two different groups of bacteria activate different genes in the mouse gut.
These genes influence a variety of processes in the intestine, such as metabolism and permeability. We identified one bacterial species in particular, Anaerostipes caccae, as the key. When we put only this species into a germ-free mouse, the mouse was protected from food allergy.
These studies show a health-promoting role for the microbiome in food allergy. It’s clear that the internal environment of the intestine is very different in infants with and without food allergy, and that this internal environment changes the biochemistry of the intestine.
Our study also suggests a way forward to harness these protective bacteria, and the molecules that they produce, as therapies to prevent and to treat food allergy. They could also work well as a diagnostic tool for predicting allergies and allergy risk. Therapies based on this idea remain 5 to 10 years away, but I am excited for their prospects. Such therapies may provide relief for children, parents, caregivers and patients living with food allergy.
By Cathryn Nagler, Bunning Food Allergy Professor, University of Chicago, President and Co-Founder of ClostraBio, Inc. This article is republished from The Conversation under a Creative Commons license. Read the original article.





Comments