In order to bioengineer complex organs, scientists need to understand the mechanisms by which those shapes are normally produced by the living organism. However, a significant knowledge gap persists between molecular genetic components identified as being necessary to produce a particular organism shape and understanding how and why that particular complex shape is generated in the correct size, shape and orientation, said the paper's senior author, Michael Levin, Ph.D., Vannevar Bush professor of biology and director of the Tufts Center for Regenerative and Developmental Biology.
An artificial intelligence system has reverse-engineered the regeneration mechanism of planaria. Credit: Daniel Lobo/Michael Levin-Tufts University
"Most regenerative models today derived from genetic experiments are arrow diagrams, showing which gene regulates which other gene. That's fine, but it doesn't tell you what the ultimate shape will be. You cannot tell if the outcome of many genetic pathway models will look like a tree, an octopus or a human," said Levin. "Most models show some necessary components for the process to happen, but not what dynamics are sufficient to produce the shape, step by step. What we need are algorithmic or constructive models, which you could follow precisely and there would be no mystery or uncertainty. You follow the recipe and out comes the shape."
Such models are required in order to know what triggers could be applied to such a system to cause regeneration of particular components, or other desired changes in shape. However, no such tools yet exist for mining the fast-growing mountain of published experimental data in regeneration and developmental biology, said the paper's first author, Daniel Lobo, Ph.D., post-doctoral fellow in the Levin lab.
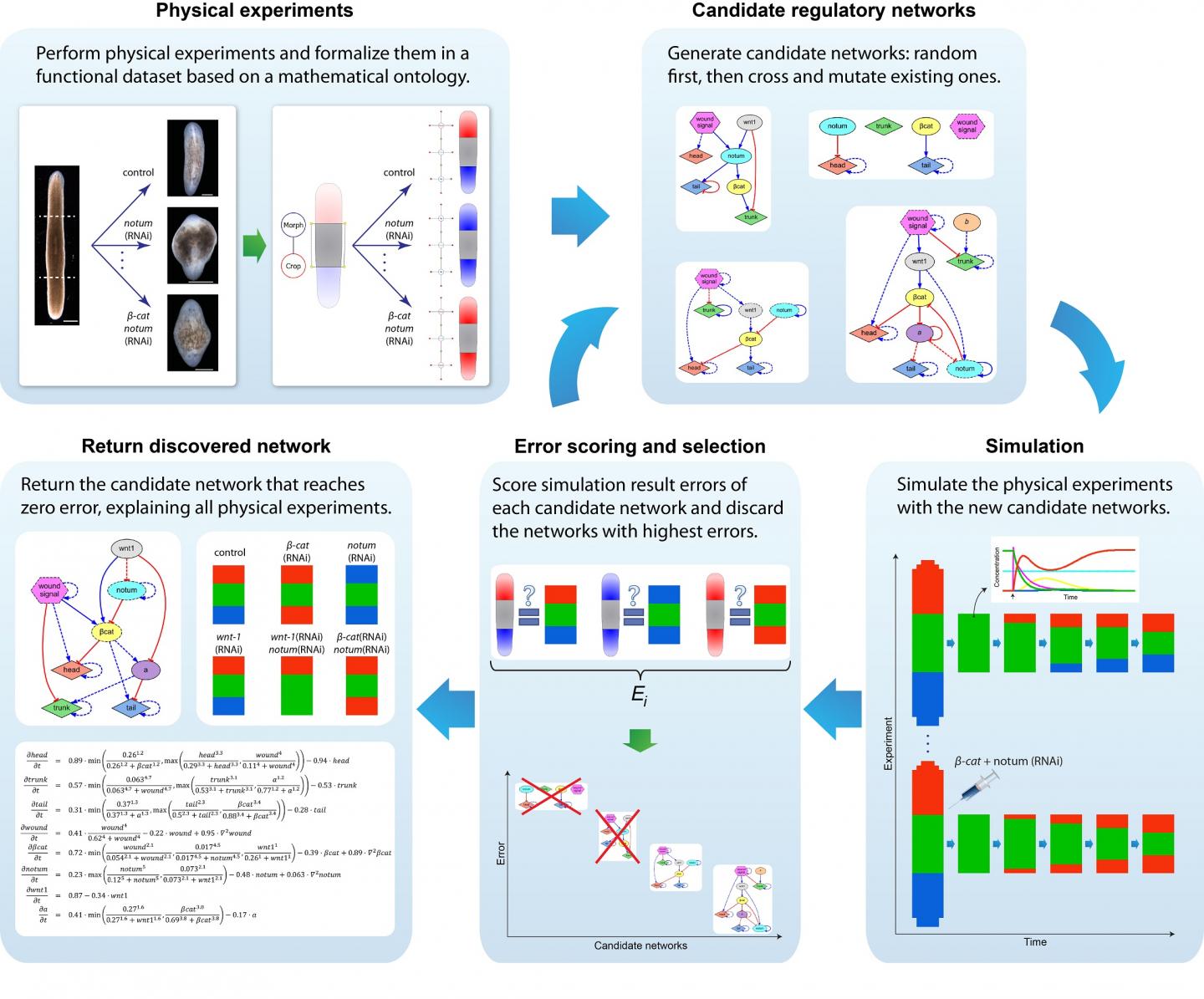
To address this challenge, Lobo and Levin developed an algorithm that would use evolutionary computation to produce regulatory networks able to "evolve" to accurately predict the results of published laboratory experiments that the researchers entered into a database.
"Our goal was to identify a regulatory network that could be executed in every cell in a virtual worm so that the head-tail patterning outcomes of simulated experiments would match the published data," Lobo said.
As expected, the initial random regulatory networks usually could not produce any of the experimental results. New candidate networks were generated by randomly combining previous networks and performing random changes, additions and deletions. Each candidate network was tested in a virtual worm, under simulated experiments. The algorithm compared the resulting shape from the simulation with real published data in the database. As evolution proceeded, gradually the new networks could explain more experiments in the database comprising most of the known planarian experimental literature regarding head vs. tail regeneration.
First Regenerative Model Discovered by Artificial Intelligence
The researchers ultimately applied the algorithm to a combined experimental dataset of 16 key planarian regeneration experiments to determine if the approach could identify a comprehensive regulatory network of planarian generation. After 42 hours, the algorithm returned the discovered regulatory network, which correctly predicted all 16 experiments in the dataset. The network comprised seven known regulatory molecules as well as two proteins that had not yet been identified in existing papers on planarian regeneration.
"This represents the most comprehensive model of planarian regeneration found to date. It is the only known model that mechanistically explains head-tail polarity determination in planaria under many different functional experiments and is the first regenerative model discovered by artificial intelligence," said Levin.
Lobo and Levin are both trained in computer science and bring an unusual perspective to the field of developmental biology. Levin majored in computer science and biology at Tufts before earning his Ph.D. in genetics. Lobo earned a Ph.D. in the field before joining the Levin lab.
The paper represents a successful application of the growing field of "robot science" - which Levin says can help human researchers by doing much more than crunch enormous datasets quickly.
"While the artificial intelligence in this project did have to do a whole lot of computations, the outcome is a theory of what the worm is doing, and coming up with theories of what's going on in nature is pretty much the most creative, intuitive aspect of the scientist's job," Levin said. "One of the most remarkable aspects of the project was that the model it found was not a hopelessly-tangled network that no human could actually understand, but a reasonably simple model that people can readily comprehend. All this suggests to me that artificial intelligence can help with every aspect of science, not only data mining but also inference of meaning of the data."
Citation: Lobo D, Levin M (2015) Inferring Regulatory Networks from Experimental Morphological Phenotypes: A Computational Method Reverse-Engineers Planarian Regeneration. PLOS Comput Biol, 11(6): e1004295. doi:10.1371/journal.pcbi.1004295. This work was supported with funding from the National Science Foundation grant EF-1124651, National Institutes of Health grant GM078484, USAMRMC grant W81XWH-10-2-0058, and the Mathers Foundation. Computation used the Extreme Science and Engineering Discovery Environment (XSEDE), which is supported by NSF grant OCI-1053575, and a cluster computer awarded by Silicon Mechanics.






Comments