Hello Again... And Bye Bye...
Hello Again... And Bye Bye...It's been a while. And now I'm back only to say goodbye. Well, not really. It's just that I've...
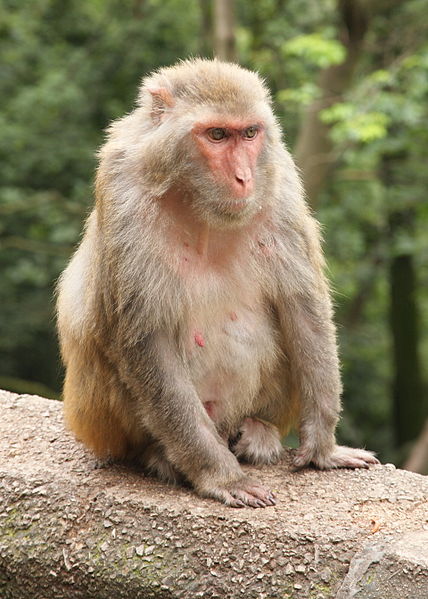 Anti-Obesity Drug?
Anti-Obesity Drug?A new compound has been shown to reduce Body Mass Index (BMI) and abdominal circumference in obese...
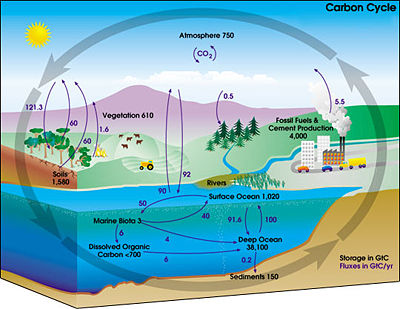 Beautiful Earth
Beautiful EarthThis video has become quite popular the last few days, so if you've already seen it, my apologies...
 The Illuminated Origin of Species
The Illuminated Origin of SpeciesTeacher turned artist Kelly Houle has set herself to the task of creating an illuminated version...