Part of the enjoyment of doing research is that ideas pop into your head all the time. Everyone has ideas, but the hard part is to choose which should be subjected to critical tests that have the primary aim of proving them wrong. That’s the most efficient way to discard bad ideas, because most of them in fact don’t work. Only after an idea survives the crucible of initial testing can it be taken more seriously and tested further. Then, if it still survives, you can publish.
Last week I described the rationale underlying a hypothesis about how life can begin, and now it is time to test it. Two older ideas are incorporated in the hypothesis, the first being that the energy of wet-dry cycles in fluctuating environments can drive the synthesis of polymers from monomers. The second is that potential organic reactants can be concentrated as orderly films on surfaces, such as the clays proposed by John Desmond Bernal in 1951.
Jim Ferris at Rensselaer Polytechnic Institute has spent much of his scientific career testing clays as potential organizing agents, and discovered that under certain conditions clay mineral surfaces can organize activated mononucleotides to produce short strands of RNA-like molecules. These are typically in the range of 10 - 15 nucleotides long, but up to 50 in length if the reaction is carefully primed. Jim’s exciting results have inspired me to test another version of an organizing surface that arises from my own research on membranes.
As I described in earlier columns, molecules like fatty acids and phospholipids can self-assemble into vesicles when dispersed in water. It is also known that when such lipids are dried, the resulting dry material is not just amorphous gunk, but instead preserves a highly ordered structure called a multilamellar array. Lamellar means that something is composed of layers.
For instance, an ordinary cake of soap is composed of layers of fatty acid molecules, and the phospholipid in egg yolk is also present as layered structures. The figure below shows what vesicles and layers actually look like.
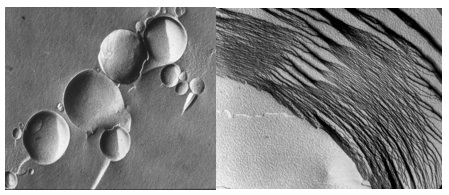
The figure on the left shows an electron micrograph of vesicles composed of lecithin, a phospholipid extracted from egg yolk, and on the right is a multilamellar array of the same lipid in a dry state.
That said, here's the hypothesis: Multilamellar structures may be able to impose order on incorporated monomers in such a way that specific polymerization reactions are promoted during condensation reactions.If so, such conditions would be expected to lead to the synthesis of long polymers.
But how could monomers get incorporated into a lipid matrix? Years ago, we discovered that when a mixture of nucleic acids and lipid vesicles is dried, the long nucleic acid molecules become trapped between the lamellae of the lipid matrix that is produced when the vesicles fuse. Furthermore, instead of being strongly adsorbed to the lipid surface, the nucleic acids return to solution as soon as water is added back to the system and then become encapsulated to produce protocells. Last week’s column showed an image of RNA that had been trapped in vesicles by this process, and the cartoon below illustrates what molecules of adenosine monophosphate might look like when they are organized between dry lipid layers.
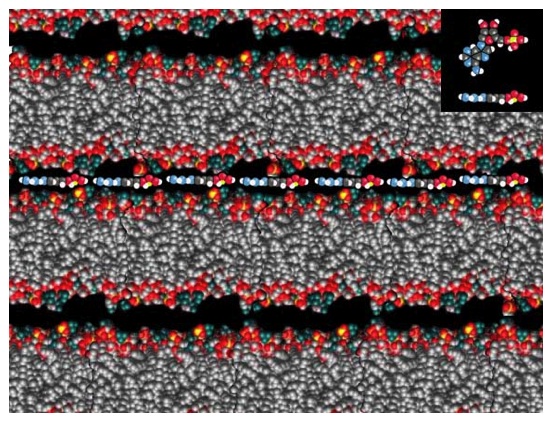

Adenosine monophosphate (AMP) is one of the four monomers of RNA, and a molecular model is shown in the inset. This figure shows how AMP could be organized in the two-dimensional space between lipid bilayers of a multilamellar array. If phosphodiester bonds can be produced between the AMP molecules, long polymeric strands of RNA will be synthesized.
Now I will describe an experimental test of the hypothesis, and some results. First, you should know something about how a university research laboratory works. The research was funded by the NASA Exobiology program which doles out grants to individual principal investigators, or PIs, in the range of $100,000 per year. This is enough to support two or three people in the lab, typically a graduate student and a post-doctoral associate. Post-docs are like interns in hospitals. They are apprentices who already have a PhD and are spending a few years learning how to be an independent investigator. Lab groups these days are often very international in composition. For the research described here, Dr. Sudha Rajamani (from India) was a post-doc who did much of the hands-on work, Dr. Felix Olasagasti (from Spain) did some of the preliminary studies, Seico Benner (from Japan) carried out several of the technical assays, and Dr. Sasha Vlassov (from Russia) did the analytical gels that finally convinced us that we were on the right track. My job was to come up with the original idea (which I first began to work on 25 years ago) and like Jack building his house, I wrote the proposal that got the grant that supplied the money that supported the team that did the experiments that were published last year.
So, what did we do? It was really very simple. We made up dispersions of lipid vesicles in very dilute hydrochloric acid (pH 3) to simulate the pH of volcanic hot springs. Mononucleotides, the monomers of RNA, were added so that there was about one nucleotide per lipid in the solution, and the solution was warmed to 90 degrees C while it was dried with a gentle stream of carbon dioxide. The gas flushed out any water molecules that were released by the reactants if they happened to form phosphodiester bonds, the basic chemical linkage of nucleic acids. The CO2 also kept out oxygen that could otherwise damage the chemicals in the reaction mix. After two hours of drying, a small amount of water was added to disperse the dried film back into vesicles and the wet-dry-wet cycle was repeated up to seven times.
This is a simulation of the conditions in the hot spring environment of a volcanic hydrothermal area on the early Earth, similar to the sites in Kamchatka and Mt. Lassen California that I described in earlier columns. We imagine that there would be a continuous drying and wetting process going on at the edges of the pools, that the water would be fairly hot (90 degrees C), weakly acidic, and that the atmosphere would have carbon dioxide but no oxygen.
So what happened? We had a wonderful surprise. When we tested the solution for the presence of polymers, we found that RNA-like molecules were present, ranging from 20 to 100 nucleotides in length. The yields were very low by the standards of organic synthesis: less than 0.1 percent of the nucleotides were linked up into longer polymers. Most researchers would ignore such low yields, but I see no particular need for high yield polymerization reactions in the prebiotic environment. In our case, even a low yield of 10 micrograms of polymer out of 10 milligrams of monomer would be equivalent to 40 trillion molecules of RNA.
The reactants on the early Earth would have been complex mixtures of hundreds, even thousands of different organic compounds, so the reactions leading to early biopolymers were almost certainly very low yields.
An important outcome of the experiment was that when the reaction was over and the last cycle of hydration was completed by adding water, the lipids formed vesicles that encapsulated the RNA products. A simple calculation shows that a few milligrams of lipid produces trillions of vesicles, each different from all the rest in its content of RNA molecules. In my view, these represent the microscopic test tubes required for the first steps toward cellular life, that is, membrane-bounded compartments containing complex mixtures of polymers that have the potential to be both catalysts (ribozymes) and carriers of information.
They are the microscopic “garbage bags” of Freeman Dyson, whom I quoted in my April 30 column.
But we can’t get too excited yet. It’s nice to find a plausible way to make polymers resembling RNA, but they are not biological RNA because the monomers are linked by 3’-5’ phosphodiester bonds (the bond of biological RNA) but also unnatural 2’-5’ bonds. This mix of bonds means that the molecules will not readily fold into a structure that can have catalytic activity like a ribozyme. Furthermore, there is no obvious way for the molecules to reproduce themselves. What would be really neat is to discover that a rare few of the strands could act as templates on which nucleotides line up, then polymerize into a second strand that has the complementary sequence of nucleotides to that of the template strand.
We have some ideas related to the fact that DNA can only contain 3’-5’ bonds, but that research is in progress so I can’t describe it just yet.
Well, this series of columns has gone on for nearly five months. If the columns are read in order start to finish, they give a brief overview of how the narrative of evolution has been extended far beyond the thin film of life in the biosphere that inspired Darwin. The origin of life, and life today, is best understood in terms of our planet and its origin, the origin of our solar system, and even the origins and deaths of stars that synthesize the atoms of life.
I am in the process of putting this narrative into a book to be published next year by the University of California Press. Now I need to get back to work on the rough draft manuscript, so instead of weekly columns I will occasionally update the progress we are making in our lab, and report interesting results from colleagues around the world. Thank you Gerhard, Patrick, Michael, and Wallace for your comments They were most appreciated. And thank you Hank Campbell for making Scientific Blogging the best on the web.






Comments