A new data analysis technique
in the journal PLoS Computational Biology improves monitoring of kidney patients and could lead to changes in the way we understand our health.
The research uses the Science 2.0 approach to make sense out of the huge number of clues about a kidney transplant patient's prognosis contained in their blood. Using big data analysis of the samples, scientists were able to crunch hundreds of thousands of variables into a single parameter indicating how a kidney transplant was faring.
That allowed the team of physicists, chemists and clinicians to predict poor function of a kidney after only two days in cases that may not previously have been detected as failing until weeks after transplant.
The extra few days would give doctors a better chance to intervene to save a transplant and improve patient recovery periods. In some cases, the team were able to predict failure from patients' blood samples taken before the transplant operation.
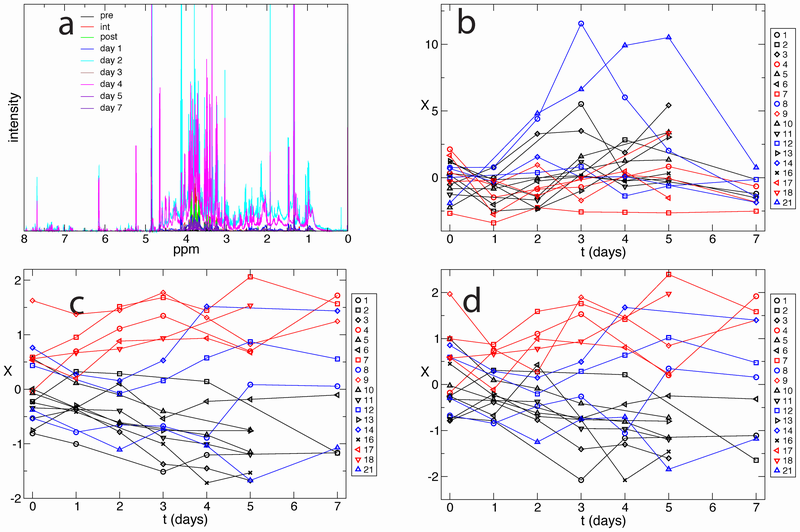
Unsupervised optimization. a) NMR spectra for blood extracts of a single patient, collected over nine time points. b) Patients trajectories projected on the first principal component do not show any feature enabling us to separate the trajectories according to the clinical assessment of the patients. The color indicates the final clinical classification of the patient: primary function patients are shown in black, delayed graft function in red, and acute rejection in blue. c) Patients trajectories projected on the optimal reaction coordinate. d) Leave-one-out cross-validation: every trajectory is projected on the optimal reaction coordinate constructed without that specific trajectory.
doi:10.1371/journal.pcbi.1003685
Dr Sergei Krivov, research fellow in the University of Leeds' Astbury Centre for Structural Molecular Biology, who led the research, said, "If you put a blood sample through Nuclear Magnetic Resonance analysis you get a very large number of different parameters that vary with the outcome for a patient.
"These are vital clues but, if you have got thousands of variables all moving in different ways in a complex system, how does a doctor bring all that information together and decide what to do? It is not possible to do this with the human mind; there are just too many variables. We have to do it with computers."
The study, which analysed daily blood samples from 18 patients immediately before and in a week-long period after kidney transplants, produced a single "optimal reaction coordinate" from the thousands of variables. This was translated to a single number (on a continuous scale from 0 to 1) describing the likelihood of a patient's state at any one time resulting in organ success or failure.
Dr Krivov said: "It is a bit like measuring GDP in the economy: a single number quantifying a huge amount of complex activity and allowing you to understand the dynamics of the system."
He added: "One of the advantages is that the output is not binary. In the past, we have tended to make decisions based on certain physical parameters. Depending on the current value or a large movement in such an indicator, we have decided whether a patient is 'healthy' or 'unhealthy' and whether or not they require treatment. At the simplest level, that could be taking their temperature. The new approach describes the dynamics of the whole system and quantifies on a continuous scale where the patient is."
Importantly, the technique does not depend on an understanding of the exact mechanism of kidney disease and is therefore, in principle, applicable in many other areas.
Dr Krivov said: "I am not a kidney specialist. I just need the data. I can then analyse it using the same equation we used here to describe the dynamics of a condition. This could be particularly powerful in areas where you are dealing with slowly developing and complex conditions, where you need to get away from a healthy-unhealthy dichotomy and engage with the incremental dynamics of the disease."
Given enough data, the technique could even be used to quantify very complex and extended processes affecting the whole population and could, ultimately, change our way of seeing our health.
"It would require a lot of data and a lot of people regularly giving their data but there is nothing in theory to stop us applying this to something like age, for instance," Krivov said.
"If you are looking at biological aging, what is the best way to quantify it? You don't just have two states: 'old' or 'young'. It is a really slow process and it is certainly not described by your passport age. If we can quantify your age in a biological way, we can change the way you see your life and health. If you have a number describing it, you can see where you are speeding up biological aging and you can then work out ways to slow it down or even reverse it."





Comments