Shaking a bag of Legos until they spontaneously form a TIE Fighter sounds absurd, yet self-assembly creates equally complex products all the time. Adding a little bit of energy to a bunch of small parts often gives them the opportunity to seek out a stable, low-energy order. We can see this in action at the breakfast table. Those last Cheerios aren’t clinging to each other for dear life—they’re seeking out a way to minimize surface tension in the milk. Whitesides at MIT has used the same principle to demonstrate self-assembly in small hexagonal tiles. Thomas Russel, Ting Xu, and their associates are doing the same thing on a molecular level.
 Self assembled monolayers, or SAMS, are a special class of self-assembled structures that form an ultrathin and highly ordered film on a surface. They are the basis of DNA microarrays, protective coatings on micromachines, and Rain-X. In nature, we see them in cells’ lipid bilayer membranes. Other researchers have been exploring the use of block-copolymer SAMS for data storage surfaces, but their efforts met with a fatal flaw—or rather, many of them. It’s difficult to pattern polymers in a dependably crystalline pattern.
Self assembled monolayers, or SAMS, are a special class of self-assembled structures that form an ultrathin and highly ordered film on a surface. They are the basis of DNA microarrays, protective coatings on micromachines, and Rain-X. In nature, we see them in cells’ lipid bilayer membranes. Other researchers have been exploring the use of block-copolymer SAMS for data storage surfaces, but their efforts met with a fatal flaw—or rather, many of them. It’s difficult to pattern polymers in a dependably crystalline pattern.The Amherst/Berkeley group overcame this by using a templated surface to guide and control the dense assembly of their polymer. They started with a sapphire wafer that had been treated to have a pronounced molecular scaled stepped pattern, a perfect seat for incoming polymer molecules. Compare this to a bowl of Cheerios with a set of ridges on one side. The Cheerios will naturally flock there, once again trying to minimize the surface tension. This first layer of Cheerios attracts more Cheerios, and soon you no longer have to chase them around the bowl with a spoon.
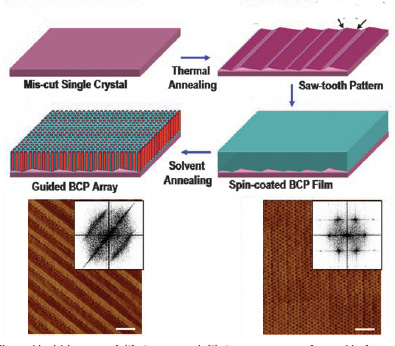 On the molecular level, this stepped surface promotes flawless arrangement of polymer molecules. Varying the step size offers further control over the density. This is important in the creation of storage devices, because a high density means that smaller features can be used to store data. The current cutting edge is about 1 terabit per square inch. A flawless square inch of this templated polymer can store ten terabits—that’s like ten of the very best iPods, and 300,000 songs.
On the molecular level, this stepped surface promotes flawless arrangement of polymer molecules. Varying the step size offers further control over the density. This is important in the creation of storage devices, because a high density means that smaller features can be used to store data. The current cutting edge is about 1 terabit per square inch. A flawless square inch of this templated polymer can store ten terabits—that’s like ten of the very best iPods, and 300,000 songs. The use of a templated surface to direct self-assembly may be novel in microelectronics, but it’s extremely reminiscent of another great self-assembled data storage unit: DNA. Understanding the templated assembly mechanism of DNA provided the foundation for polymerase chain reaction, or PCR. Vastly simplified, PCR uses a single helix of DNA as its patterned surface, attracting the assembly of a new chain of complementary nucleotides. When a complete double helix is assembled, it is split, yielding two single-stranded DNA helices—two patterned surfaces to attract two new sets of nucleotides. Each cycle of PCR provides new templated strands, and new DNA, until enough copies have been made to perform experiments.
This DNA cloning technique enabled huge advances in genetics, and was instrumental in the human genome project. Not bad for a bunch of lazy molecules.
Soojin Park, Dong Hyun Lee, Ji Xu, Bokyung Kim, Sung Woo Hong, Unyong Jeong, Ting Xu, Thomas P. Russell. Macroscopic 10-Terabit–per–Square-Inch Arrays from Block Copolymers with Lateral Order. Science 323, 1030 (2009).
N. Bowden, S. R. J. Oliver, G. M. Whitesides. Mesoscale Self-Assembly:Capillary Bonds and Negative Menisci. J. Phys. Chem. B, 2000, 12.





Comments