The fast changing pace of these secreted proteins also suggests that they are the basis for bacterial adaptation, and should be the first stop for researchers looking for new ways to fight bacterial infection, exploit their resources or simply trying to understand these organisms better.
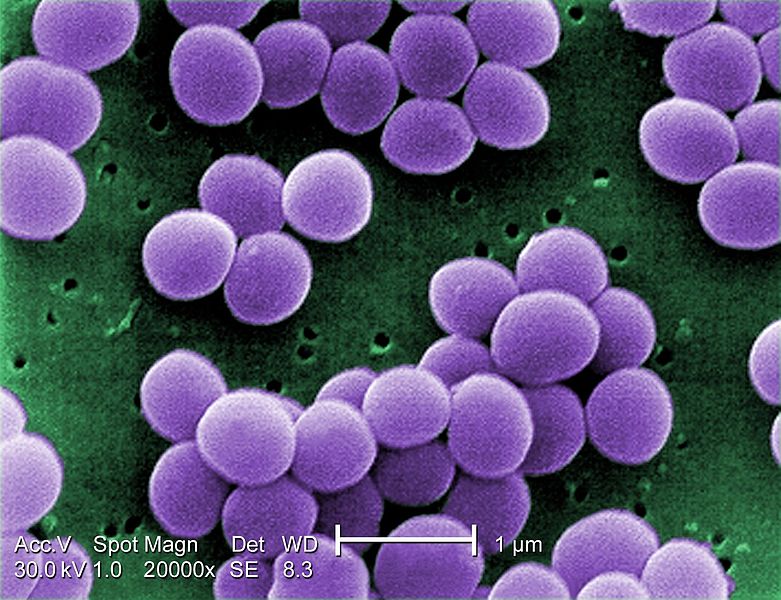
Scanning electron micrograph of an antibiotic resistant Staphylococcus
bacteria by Janice
Haney Carr with the higher
resolution of transmission electron microscopy would be seen the thickening of
the cell wall that is responsible for the reduced susceptibility to antibiotics
It has become clear, that though we can be killed by bacteria we cannot live without them either. From stimulating our immune system, provide vitamins that we cannot synthesize, affect diabetes and obesity to even protect us against other pathogens, bacteria can be both for or against us. The increasing awareness of this always-present relationship has turned more urgent than ever the need to understand better these microorganisms so we could tip the balance in our favor when necessary.
And the core of any bacterium's life is the secretome – bacteria’s secreted proteins that mediate their relationship with the outside world and have functions as diverse as capturing food, changing the environment to suit bacteria’s needs or interacting with other organisms. What is particularly interesting with this approach to the outside is that is has a crucial problem – once secreted these proteins become “public goods” available to be exploited by any organism in the vicinity. And if the “freeloaders” cannot be stopped, the bacteria producing proteins can be driven to extinction and replaced by the cheaters.
A strategy to avoid this - seen for example in Escherichia coli a common bacterium in humans – is to make the secretome genes highly mobile using horizontal gene transfer, a non-sexual process where genes “jump” from one individual to another. As the secretome genes (also called social genes due to their role in bacteria-bacteria interaction) spread to other bacteria, the population producing proteins increases reducing freeloaders’ numbers until their stealing no longer matters.
But genes with high mobility also have high chances of being lost or changed as they leave a genome to enter other. And this - together with the evolutionary competition (where a species evolve as result of having to escape or pursue other) occurring between bacteria and their hosts, parasites or predators – made researchers Teresa Nogueira, Marie Touchon and Eduardo P. C. Rocha think that social genes should be under strong evolutionary pressure to change rapidly and, as such, vital in bacteria’s evolution.
To test this hypothesis they looked at more than 400 bacterial genomes, many pathogenic for humans, first grouping them according to their genetic similarities and common ancestors, to then compare their genes according to the localization (inside and outside of the cell) of their proteins. They found that although only a small part of the bacterial genome produced secreted proteins, its size was relatively constant and not dependent of the bacterial genome size with this consistency suggesting an important role for the organism.
Next, thry searched for the social genes within the genome and, confirming
their suspicions, these were located in its most mobile parts: the accessory genome (a supernumerary part
of the genome where there are no genes essential for survival) and the plasmid (a piece of extra DNA outside
of the chromosomes).
Not only that but they also found that social genes existed
only in some bacteria species within a group indicating they had appeared
recently (otherwise they would have had time “jump” into the other species), supporting
the idea that these were genes changing rapidly. But
were they changing faster than the rest of the genome?
To answer that the researchers looked for orthologous proteins (proteins in different species that originated from a common ancestor), comparing their rate of change in the secretome with the rest of the genome. Again, not only secreted proteins had higher rates of change, but they also showed a type of change, called non-synonymous substitutions, that is known to lead to proteins with new biological functions.
In conclusion, social/secretome genes
not only are localized in the most mobile parts of the genome, but they also show
signs of accelerated change and of having evolved to produce new proteins recently
supporting the idea that they are crucial in bacterial adaptation.
“Remarkably what our work shows,” says Nogueira, “is that cell localization shapes genome evolution.
Bacteria evolution is triggered by the interaction with neighboring bacteria,
by what they are producing and releasing to the surroundings (the secretome). Most
importantly, this also means that the repertoire of genes coding for secretome
can be seen as a therapeutic target for the control of bacterial infection. And
with resistance to antibiotics already considered one of the greatest threats
of modern health, all help is welcome.”
Journal article
Nogueira T, Touchon M, Rocha EPC, Rapid evolution of the sequences and gene repertoires of secreted proteins in bacteria, (2012) PLoS One 7(11): e49403. doi:10.1371/journal. pone.0049403 http://www.plosone.org/article/info%3Adoi%2F10.1371%2Fjournal.pone.0049403






Comments