How does a stem cell decide what specialized identity to adopt – or simply to remain a stem cell? A new study suggests that the conventional view, which assumes that cells are “instructed” to progress along prescribed signaling pathways, is too simplistic. Instead, it supports the idea that cells differentiate through the collective behavior of multiple genes in a network that ultimately leads to just a few endpoints – just as a marble on a hilltop can travel a nearly infinite number of downward paths, only to arrive in the same valley.
The findings, published in the May 22 issue of Nature, give a glimpse into how that collective behavior works, and show that cell populations maintain a built-in variability that nature can harness for change under the right conditions. The findings also help explain why the process of differentiating stem cells into specific lineages in the laboratory has been highly inefficient.
Led by Sui Huang, MD, PhD, a Visiting Associate Professor in the Children’s Hospital Boston Vascular Biology Program (now also on the faculty of the University of Calgary), and Hannah Chang, an MD/PhD student in Children’s Vascular Biology Program, the researchers examined how blood stem cells “decide” to become white blood cell progenitors or red blood cell progenitors.
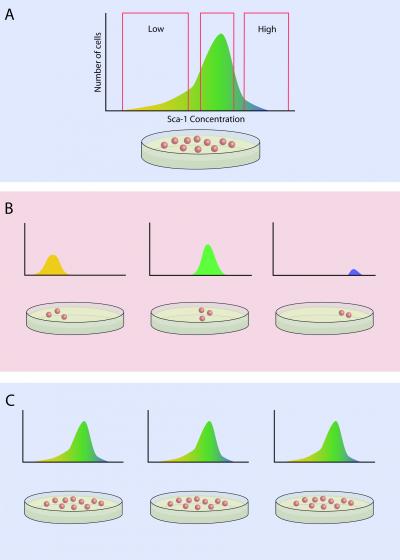
They began by examining populations of seemingly identical blood stem cells, and found that a cell marker of “stemness,” a protein called Sca-1, was actually present in highly variable amounts from cell to cell – in fact, they found a 1,000-fold range. One might think that low Sca-1 cells are simply those cells that have spontaneously differentiated. However, when Huang and Chang divided the cells expressing low, medium and high levels of Sca-1 and cultured them, each descendent cell population recapitulated the same broad range of Sca-1 levels over nine days or more, regardless of what levels they started with.
“We then asked, are these cells also biologically different?” says Huang, the paper’s senior author. “And it turned out they were dramatically different in differentiation.”
Blood stem cells with low levels of Sca-1 differentiated into red blood cell progenitors seven times more often than cells high in Sca-1 when exposed to erythropoietin, a growth factor that promotes red blood cell production. Conversely, when stem cells were exposed to granulocyte–macrophage colony-stimulating factor, which stimulates white blood cell formation, those that were highest in Sca-1 were the most likely to become white cells. Yet, in both experiments, all three groups of cells retained characteristics of stem cells.
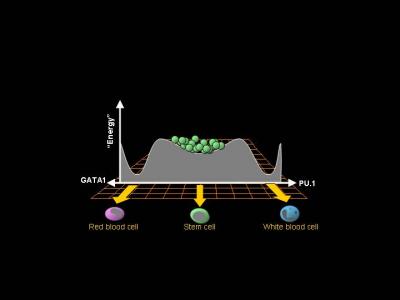
Huang and Chang then looked at the proteins GATA1 and PU.1, transcription factors that normally favor differentiation into red and white blood cells, respectively. Blood stem cells that were low in Sca-1 (and most prone to become red blood cells) had much more GATA1 than did the high- and medium-Sca-1 cells. Stem cells high in Sca-1 (and least prone to become red blood cells) had the highest levels of PU.1.
But most important, the differences in Sca-1, GATA1 and PU.1 levels across the three cell groups became less pronounced over time, as did the variability in the cells’ propensity to differentiate, suggesting that the differences are transient.
In a final step, Huang and Chang used microarrays to look at the cells’ entire genome. Again, they found tremendous variability within the apparently uniform cell population: more than 3,900 genes were differentially expressed (turned “on” or “off”) between the low- and high-Sca-1 cells. And again, this variability was dynamic: the differences diminished over time, with gene activity in both the low- and high-Sca-1 cells becoming more like that in the middle group.
Together, the findings make the case that a slow fluctuation or cycling of gene activity tends to maintain cells in a stable state, while also priming them to differentiate when conditions are right.
“Even if cells are officially genetically identical and belong to the same clone, individual members of that population are quite different at any given time,” says Huang. “This heterogeneity has usually been seen as random ‘measurement noise,’ and, more recently, as ‘gene expression noise.’ But it turns out to be very important, and is the basis for stem cells’ multipotency – their ability to differentiate into multiple lineages.”
“Nature has created an incredibly elegant and simple way of creating variability, and maintaining it at a steady level, enabling cells to respond to changes in their environment in a systematic, controlled way,” adds Chang, first author on the paper.
Practically speaking, the work suggests that stem cell biologists may need to change their approach to differentiating stem cells in the laboratory for therapeutic applications.
“So far the process has been highly inefficient – only 10 to 50 percent of cells respond to the hormone or whatever is given to make them differentiate,” Huang says. “That is because of the cells’ inherent heterogeneity. People have been finding more and more sophisticated stimulator cocktails, but we could make the process more efficient by harnessing the heterogeneity and identifying cells that are already highly poised to differentiate.”
Chang has already done follow-up experiments showing that stem cell differentiation can be made dramatically more efficient by choosing the right subpopulation of stem cells and stimulating them promptly, while they are most apt to differentiate. “I’m not doing anything complicated – just using what nature already has,” she says.
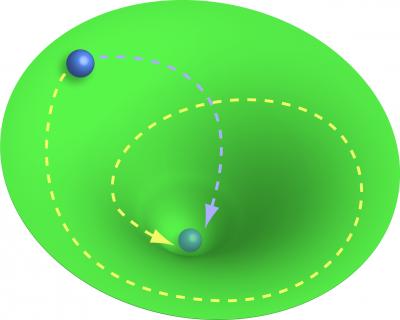
But the findings also challenge biologists to change how they think about biological processes. The work supports the idea of biological systems moving toward a stable “attractor state,” a concept borrowed from physics. In this case, blood stem cells tend to remain blood stem cells, yet they experience inherent fluctuations in gene activity and protein production that can sometimes be enough to tip the balance and cause them to fall into other attractor states – namely, red or white blood cell progenitors. Specific growth factors can tip the balance, but these factors are part of an overall landscape that guides cells toward different destinies. A marble going downhill will eventually end up in a valley, but which valley it falls into depends on the shape of the landscape.
“Growth or differentiation factors merely increases the probability that a cell will grow or differentiate,” says Donald Ingber, MD, PhD, a co-author on the paper who, with Huang, served as Chang’s mentor on the project. “Cell differentiation is an ensemble property, a collective behavior, inherent in the system’s architecture and set of regulatory interactions.”
A previous study by Huang established, for the first time, that a given cell can exhibit a very different pattern of gene activity from its neighbor, taking a very different path through the landscape, yet end up in the same valley. He and his colleagues exposed precursor cells to two completely different drugs (DMSO and retinoic acid) and closely monitored the cells’ gene expression. Both groups of cells eventually differentiated to become neutrophils (a type of white blood cell), but the molecular paths they took and their patterns of gene expression were completely different until day seven, when they finally converged.
The landscape analogy and collective “decision-making” are concepts unfamiliar to biologists, who have tended to focus on single genes acting in linear pathways. This made the work initially difficult to publish, notes Huang. “It’s hard for biologists to move from thinking about single pathways to thinking about a landscape, which is the mathematical manifestation of the entirety of all the possible pathways,” he says. “A single pathway is not a good way to understand a whole process. Our goal has been to understand the driving force behind it.”
This study was funded by the Air Force Office of Scientific Research, the National Institutes of Health, the Presidential Scholarship, the Ashford Fellowship of Harvard University, and the Army Research Office.





Comments