A cool movie of a cell crawling. The red bursts at its front are a fluorescent protein that localizes to growing pseudopods.
To hunt down your prey you have to be able to sense it, of course, but you also have to be pretty good at going straight. Even the best hunter will lose the trail every once in a while, but odds are he can pick up the trail again if he keeps going straight for a few more paces. But if you spun him around until dizzy and gave him a few shots of tequila, he might not do so well at staying on target. Migrating cells will often lose track of their prey too, since their microscopic environments are very noisy (technical term). If cells want to eventually find their target, then, they need to be able to keep going straight and not get too distracted by the mess. So how do these cells stay on target even when they lose the trail? This is the question asked by a new paper out in PLoS One (Cooper, Wingreen, Cox, 2012).
Cells are actually pretty good at going straight, even for several minutes (Li, Norrelykke, Cox, 2008; Bosgraaf&van Haastert, 2009). But how do they do this? The control circuits that make cells move are very complex, with lots of redundant pathways, and there’s still a lot we don’t know about them. So the approach taken by this new study was to step back and look at the larger picture of what we do know. We know how different larger modules of the system behave, so by putting them together we might be able to figure out how the whole cell-crawling system keeps going straight, even without knowing what every single molecule does.
So what motility modules do the researchers use in their broader approach? First, they observe that the pseudopods used by cells to crawl around behave like excitable systems. A pseudopod just means a part of the cell that pushes out and grabs onto the ground in front of it, which the cell then uses to pull itself forward. And to understand an “excitable system,” walk over to a somewhat stressed coworker and gently poke him or her. There will probably be little response. Now repeat until finally the coworker explodes, screams his head off, and then finally slumps back down into his chair, exhausted. Congratulations, you just provoked an excitable burst of activity! Your coworker has some low ground state with no screaming, but given some random amount of stimulation he switches into screaming mode. Then after he’s done, he enters a third state, where he’s so tired that he has to recover for a little while before he can burst again.
Pseudopods in cells behave like your excitable coworker – they randomly erupt in bursts of pushing out, but then they stop and the local cell membrane takes a little while to recover before starting again. So excitable pseudopods are the first motility module. The study’s second motility module is “global repression”: one burst of activity seems to repress new pseudopods from forming anywhere else in the cell. Only one excitable coworker is screaming at a time, the other ones wait their turn.
These motility modules were supported by previous observations, but the new hypothesis in this study was that after a burst of pseudopod activity, that local area of the cell membrane is then even more excitable for future pseudopods. Once your coworker recovers, he’s even more on edge than he was before, so it takes fewer pokes to get him to explode again.
This increased excitability gives the cell a way to remember which way it’s going. If it’s been heading North for a while but then randomly veers off to the West a bit (because in biology things are just messy like that), then, since its North side is still more excitable, odds are that the next pseudopod will turn it back North again.
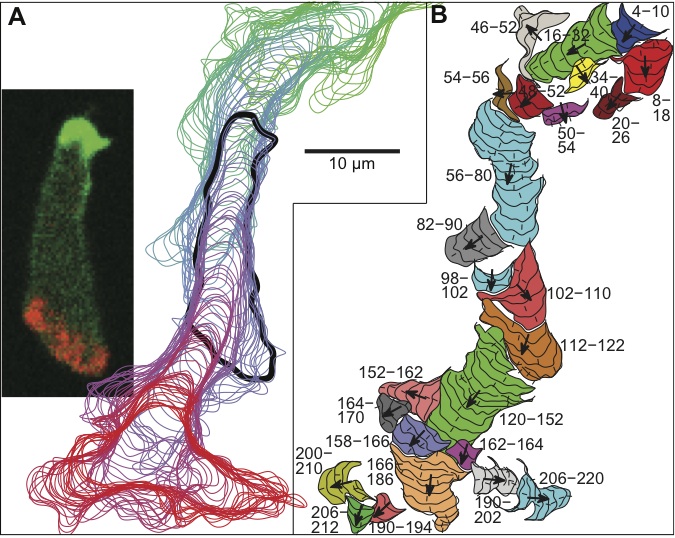
So how did the scientists test this idea? First, they made a computer model and did some simulations to show that model cells do indeed move like real cells. Then, they looked at some more detailed predictions from the model, things nobody else had looked for yet. For example, the local increase in excitability should only last for a limited time, so if you pick out only the pseudopods that happen to have a long delay between them, you should see the directional memory start to disappear. Their model made four specific predictions about similar details of pseudopod dynamics, and when they looked at real cells, experimental data confirmed all four of the predictions.
The cool thing about this model is that it gives scientists a way to understand how the things they already know about cell motion can add up to help cells go straight, even without knowing all the details. Scientists already knew that pseudopods behave like excitable systems (module 1), and there’s usually only one excitable burst at a time (module 2). Just by assuming a temporary memory of where there was a previous pseudopod (module 3) and wiring them all together, we can understand directed motion from this broader perspective. By now knowing to look for this third module, the directional memory effect, scientists can now study its details and home in on what genes and proteins are behind it.
The hope is that this framework could one day help doctors prevent inflammatory cells from moving in where they don’t belong, or even keep cancer cells from escaping their main tumor and invading healthy tissue. Of course that will require a lot more work, but I still think this paper is really cool. Plus I hear its first author is uncannily dashing in a lab coat. Ok, the other authors too (cough cough, thesis committee, cough cough).
------------------------
Cooper RM, Wingreen NS, Cox EC (2012) An Excitable Cortex and Memory Model Successfully Predicts New Pseudopod Dynamics. PLoS ONE 7(3): e33528. doi:10.1371/journal.pone.0033528





Comments