Researchers at Columbia University Medical Center have illuminated a window into how abnormalities in microRNAs, a family of molecules that regulate expression of numerous genes, may contribute to the behavioral and neuronal deficits associated with schizophrenia and possibly other brain disorders.
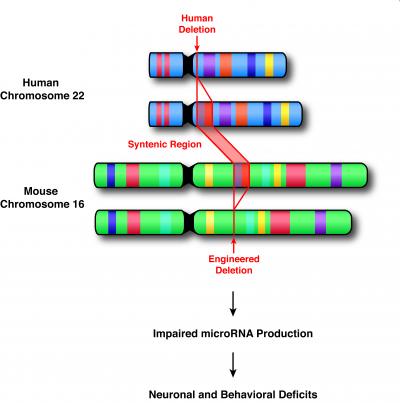
Maria Karayiorgou, M.D., professor of psychiatry, and Joseph A. Gogos, M.D., Ph.D., associate professor of physiology and neuroscience at Columbia University Medical Center explain how they uncovered a previously unknown alteration in the production of microRNAs of a mouse modeled to have the same chromosome 22q11.2 deletions previously identified in humans with schizophrenia.
“We’ve known for some time that individuals with 22q11.2 microdeletions are at high risk of developing schizophrenia,” said Karayiorgou, who was instrumental in identifying deletions of 22q11.2 as a primary risk factor for schizophrenia in humans several years earlier. “By digging further into this chromosome, we have been able to see at the gene expression level that abnormalities in microRNAs can be linked to the behavioral and cognitive deficits associated with the disease.”
The investigators modeled mice to have the same genetic deletion as the one observed in some individuals with schizophrenia and examined what happens in the expression of over 30,000 genes in specific areas of the brain. When they discovered that the gene family of microRNAs was affected, they suspected that the Dgcr8 gene was responsible.
The Dgcr8 gene is one of the 27 included in the 22q11.2 microdeletion and has a critical role in microRNA production, so this was a logical hypothesis. Indeed, when they produced a mouse deficient for the Dgcr8 gene, and tested it on a variety of cognitive, behavioral and neuroanatomical tests, they observed the same deficits often observed in people with schizophrenia.
“Our studies show that alterations in microRNA processing result in synaptic and behavioral deficits,” said Dr. Gogos. Drs. Karayiorgou and Gogos have partnered together to decipher the role of individual genes from 22q11 in the development of schizophrenia by using human genetics and animal model approaches.
The significance of this work is that it implicates a completely novel, previously unsuspected group of susceptibility genes and brings investigators a step closer to understanding the biological mechanisms of this disorder. Implication of such a large family of genes (the most recent estimate puts the number of human microRNAs at at least 400 that influence the expression of as many as a third of all genes) could partly account for the genetic complexity associated with this devastating disorder and explain some of the difficulties that the researchers have encountered in their efforts to pinpoint individual genes.
“Our hope is that the more we know about the genes involved in schizophrenia, the more targeted treatment can be,” said Dr. Gogos.
“Much in the way that cancer patients who have tested for a particular gene, such as BRAC1, can be tested and then treated with protocols designed specifically for them, we want to be able to know enough about the schizophrenic brain to target treatments to individual patients.”
The next step for the researchers is to find the many genes whose expression is controlled by the identified deficient microRNAs, which could in turn be involved in the pathogenesis of schizophrenia. Much more study and identification of other genetic variants must be done to further illuminate the disease’s genetic underpinnings, according to Drs. Karayiorgou and Gogos.
In this study, in addition to their colleagues from Columbia, first authors Kimberly Stark and Bin Xu, as well as co-authors Wen-Sung Lai, Ruby Hsu and Hui Liu, Drs. Karayiorgou and Gogos collaborated with Anindya Bagchi and Alea Mills at Cold Spring Harbor Laboratory and Xiang Wan and Paul Pavlidis at the University of British Columbia.
This research was supported by the National Institutes of Health, the Lieber Center for Schizophrenia Research at Columbia University Medical Center, New York Academy of Sciences, a McKnight Brain Disorders Award, the National Alliance for Research on Schizophrenia and Depression, and the EJLB Foundation.






Comments